5 Tips for Setting Up Your PCR
Script
Here are five basic tips for setting up your PCR.
First, make sure to choose a polymerase that matches your needs. If you are unsure which polymerase will meet your specific needs, please consult the DNA polymerase selection chart at ConfidentPCR.com.
Second, take time to design your primers carefully. This will help reduce the amount of time you spend troubleshooting. Primers should typically be between 20 and 30 nucleotides in length and between 40 to 60% GC content. You also want to be sure to have your primers with Tms that are within five degrees of each other.
Third, calculate your annealing temperature using the TM calculator on neb.com. Not all enzymes are alike and buffer components can play a critical role in determining the annealing temperature. If you're switching to a new enzyme, you should always recalculate the annealing temperature even if you've used these primers with another enzyme in the past. The TM calculator on the neb.com website takes into account the specific composition of the buffer when determining the annealing temperature.
Fourth, when possible, calculate the GC content of your target. For GC-rich amplicons, use a polymerase specifically designed for GC-rich PCR, like OneTaq or Q5 DNA polymerases. If high GC additives are included with the polymerase, please follow our usage recommendations for the best results.
Fifth, set up reactions on ice using chilled components and add the reactions to a thermocycler that's been pre-heated to your denaturation temperature. Alternatively, try a hot start enzyme such as our Q5 or OneTaq hot start DNA polymerases for convenient room temperature setup and the additional benefit of increased specificity.
Related Videos
-
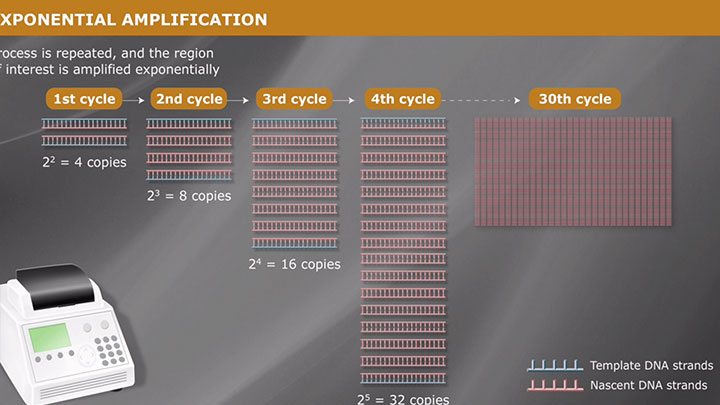
Overview of PCR -

Tips for using the Monarch® PCR & DNA Cleanup Kit -
Choose the Right DNA Polymerase for PCR

