Identification and Characterization of Protein Glycosylation
Script
The overall goal of the following experiment is to illustrate the use of glycosidases for the enzymatic deglycosylation and analysis of N and O-linked glycosylation on a model glycoprotein.
This is achieved by treating a glycoprotein with PNGase F or with the Protein Deglycosylation Mix. Also, the Protein Deglycosylation Mix was supplemented with a mixture of additional exoglycosidases, which sometimes help remove otherwise resistant sugars. We illustrate this method using a model glycoprotein, recombinant human chorionic gonadotropic β or hCG β.
Next, SDS-PAGE followed by Coomassie blue staining and sugar-specific staining method, Pro-Q Emerald 300, are used in order to analyze the deglycosylated hCG β sample. Results are obtained which show that hCG β is heterogeneously glycosylated and contains multiple glyco-forms based on the differences in protein migration with and without glycosidase treatment and the diminished signal on Pro-Q staining, as the glycans are successively removed.
Alicia Bielik:
The main advantage of this technique over existing methods, such as mass spectrometry analysis, is that it's simple, uses common laboratory equipment, and is good for the novice glycobiologist. This method can answer key questions in the glycobiology field such as, "Is my protein glycosylated?"
Anthony Sheets:
To begin enzymatic deglycosylation, thaw the supplied buffers from the protein deglycosylation kit. Mix the tubes thoroughly and keep at room temperature. Place the enzyme containing vials on ice. Dissolve the contents of the hCG β vial in 600 microliters of distilled water and keep on ice. After preparing 1 milliliter of 1x G7 buffer by diluting the 10x stock in distilled water, use 25 microliters to dilute 0.5 microliters of PNGase F. Prepare the exoglycosidase mix by combining 2 microliters each of the four exoglycosidases. Next, add hCG β and 10x glycoprotein denaturing buffer to 7 numbered PCR tubes, as indicated in the written protocol. Cap the tubes and mix gently.
Denature the proteins in the thermocycler by incubating 10 minutes at 94° Celsius, followed by a 4° Celsius hold. After removing the tubes from the thermocycler, centrifuge to remove any visible condensation. To each reaction tube, add the remaining reagents according to the written protocol. Close the PCR tubes using new caps, then mix the tubes by gently tapping four times. After spinning down the contents, place the tubes in the thermocycler. Incubate at 37° Celsius for 4 hours. Follow by cooling the samples to 4° Celsius.
To set up SDS-Page, prepare fresh 3x reducing SDS-loading buffer with DTT. Add 12.5 microliters of the prepared 3x reducing SDS-loading buffer to each sample. Close the tubes with new caps and gently tap the tubes to mix. Incubate the tubes in the thermocycler at 94° Celsius for 5 minutes and then cool to 4° Celsius. Following incubation, load 30 microliters of each sample and 10 microliters of the protein marker on a 10-20% tris-glycine gel, saving the remainder of each sample. Electrophorese the gel at 130V until the dye front is near the bottom of the gel. When the gel has finished running, remove the gel from the cast and place it in a small, plastic box with enough Coomassie blue stain to cover the gel. Stain the gel for one hour with gentle agitation.
After the gel is stained, wash three times for 30 minutes in 50 milliliters of destained solution. Record the images using a white light transilluminator or scanner. Alternatively, the gel can be dried between sheets of cellophane in a frame.
Run the remainder from each sample and a protein marker on a 10-20% tris-glycine gel. While the gel is running, dissolve the Pro-Q Emerald reagent with DMF and prepare the stock-fix, wash, and oxidizing solutions for the stain, following the product manual provided with the kit. When the electrophoresis is complete, remove the gel from the cast and place it in a plastic box. Fix the gel by adding 100 milliliters of the prepared fix solution. Leave the gel overnight at room temperature with gentle agitation. The following day, wash the gel with 100 milliliters of the prepared wash solution for 10-20 minutes at room temperature with gentle agitation, repeating the wash a second time with fresh wash solution.
Next, oxidize the carbohydrates by incubating the gel with gentle agitation for 30 minutes in 25 milliliters of the prepared oxidizing solution. Follow the incubation with two additional washes. While the gel is washing, prepare fresh Pro-Q Emerald 300 stain by adding 500 microliters of the Pro-Q Emerald 300 reagent solution to 25 milliliters of staining buffer provided in the kit. Stain the gel by incubating in the prepared stain under dark condition with gentle agitation for 90-120 minutes. Subsequent to gel staining, repeat the two wash steps as described earlier. Then, record the images with a UV transilluminator at 300 nanometers. As a final step, compare the images of the Coomassie-stained gel with the Pro-Q Emerald-stained gel.
The changes in protein migration after enzymatic deglycosylation can be seen when the control sample is compared with the PNGase F treatment to remove N-glycans and to the deglycosylation-mixed treatment to remove N and O glycans. No further reduction in size is seen after digesting with additional glycosidases. Besides a change in mass, bands become sharper as glycans are removed. A band running under the 17 kilodalton marker probably represents the fully deglycosylated hCG β polypeptide. Lanes five to seven show the bands corresponding to the glycosidases. Other bands might derive from incomplete deglycosylation or from multiple, unidentified proteins present in the hCG β sample. The Emerald green reagent oxidizes and stains all gylcans present in a protein molecule, therefore, the intensity of the signal decreases as hCG β is enzymatically deglycosylated.
The residual signal in lanes three and four indicate the presence of glycan motifs, which are resistant to the enzymes used. The additional glycosidases used in lane four remove a few extra sugar residues. The protein migration is the same, but a slight reduction in intensity in staining can be seen. Resistant sugar moieties were not present in all protein species. Some bands were not detected by Emerald green, indicating that they were extensively deglycosylated. Additional data support the conclusion that hCG β is heterogeneously glycosylated. The lower band on lane two is faint on the Emerald green image, while the upper band on lane two is bright, indicating that many glycan groups are still present. These data support the conclusion that recombinant hCG β expressed in mouse cells contains multiple glycoforms due to the inherent heterogeneity of glycosylation.
Alicia Bielik:
Following this procedure, other methods, such as mass spectrometry, can be performed in order to answer additional questions, such as, what is the rate of occupancy, what is the extent of glycosylation, or what is the fine structure of the glycans. After watching this video, you should have a good understanding of how to use glycosidases for the enzymatic deglycosylation and analysis of N and O-linked glycans on glycoproteins.
The choice of detection can be difficult because protein-staining regions are useful only if the deglycosylation results in a significant shift in molecular mass. For other proteins, we have seen abnormal migrations after deglycosylation and have found that any change in migration is evidence that the protein has been deglycosylated.
Related Videos
-
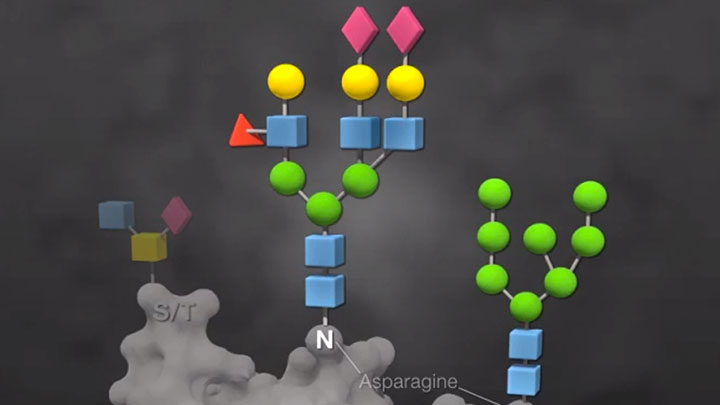
Overview of Glycobiology -

One-step protocol for deglycosylation with Rapid PNGase F -

Two-step protocol for deglycosylation with Rapid PNGase F

