Cloning With Restriction Enzymes
Script
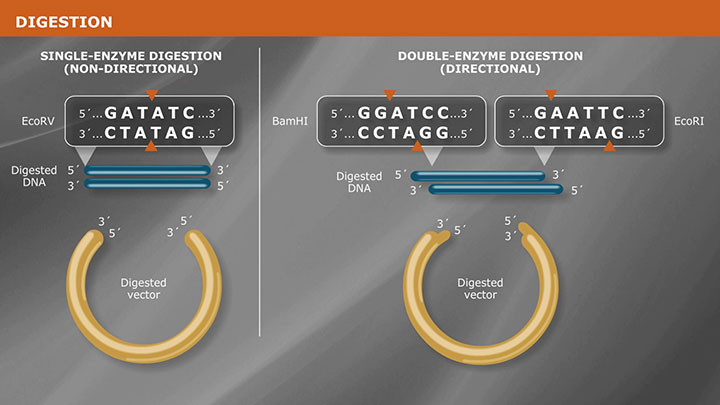
Restriction enzymes identify a specific recognition sequence in the DNA, and generate double-stranded breaks in the DNA duplex. Cleavage results in fragments with either a 5' or 3' overhang, commonly referred to as sticky ends, or no overhang, referred to as blunt ends.
The 5' phosphates and 3' hydroxyls are maintained after cleavage, leaving the DNA ends ready to be joined via ligation.
The ability of some restriction enzymes to predictably cleave DNA and generate distinct DNA bands with ligatable ends has made them an invaluable tool for recombinant DNA technologies.
When selecting restriction enzymes for use in a cloning experiment, it is important to: determine which enzymes will produce ends compatible with the selected vector confirm that recognition sites do not occur within the DNA fragment to be cloned and examine the methylation sensitivity of your selected enzyme to confirm that host methylation by dam and/or dcm methylases will not block cleavage.
NEB offers a variety of online interactive tools to aid in selecting which enzyme to use, as well as parameters for setting up reactions. Visit CLONEWITHNEB.com for a full list of products available for this application.
Related Videos
-
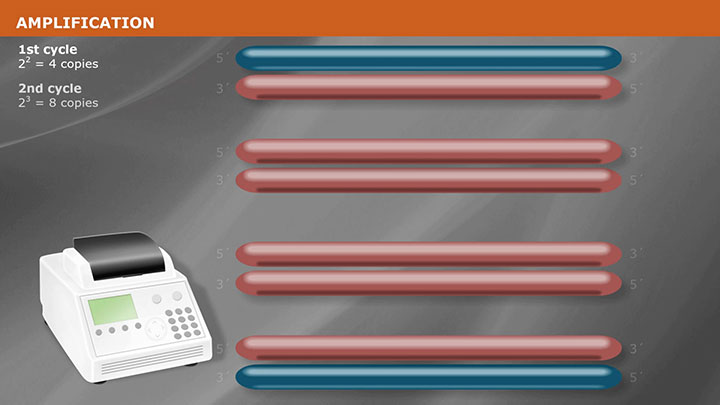

Overview of PCR Cloning -
NEB® TV Ep. 8 – Cloning and DNA Assembly -
Overview of Traditional Cloning

