NEB® TV Ep. 8 – Cloning and DNA Assembly
Script
NEBTV, what's trending in science.
Deana Martin:
Welcome to NEBTV. Today I'm joined by NEB scientists Chris Taron and Lisa Maduzia. Hey guys.
Lisa Maduzia:
Hi Deana.
Chris Taron:
Hey Deana.
Deana Martin:
And we're talking about cloning and DNA assembly.
In our Science in "60", Chris and Lisa will tell us about why they prefer traditional cloning or DNA assembly methods. Then we're going to learn a little bit about Golden Gate technology. Then we'll learn about some online tools that NEB offers to support Golden Gate and DNA assembly. And finally, we'll talk to one of our product managers to hear about a recent tech support question that they received on DNA assembly methods.
You guys ready?
Chris Taron:
Yes.
Lisa Maduzia:
Hey, let's do it.
Deana Martin:
Great.
Chris Taron:
My lab expresses proteins and yeasts. This involves cloning a target gene into a yeast expression vector in E. Coli using traditional cloning methods and takes about four days.
With NEBuilder HiFi DNA Assembly, we can entirely bypass E. Coli cloning steps.
The same linear expression cassette can be assembled in vitro in just one afternoon. Additionally, we can make multiple expression cassettes in parallel using different arrangements of promoters selectable markers and signal peptides. The result is faster and more flexible strain construction.
The method is also great even for simple cloning experiments.
Lisa Maduzia:
That's great, but I still clone using traditional methods. I typically only need to clone one or two fragments. Restriction enzyme cloning is easy, it's efficient, I know how to do it and I have all the components readily available right in my freezer.
It works for what I'm doing so I don't need to switch to other methods. There are all these great tools available to help me along with my experiments. NEB Cloner, NEB BioCalculator, Enzyme Finder and the one we all know and love, the NEB catalog.
Golden Gate Assembly presentation:
Golden Gate assembly is a method for assembling multiple DNA fragments relying on the activity of Type IIS endonucleases that cleave outside of non-palindromic recognition sites.
In this example, the destination vector is designed to include BsaI recognition sequences in opposing directions to allow for the assembly of the desired fragments.
Using PCR BsaI, recognition sites are incorporated into the vector and fragments in the proper orientation. Following amplification, the linearized vector and PCR amplified fragments are ready to be assembled in a single tube reaction containing BsaI and T4 DNA ligase.
Cleavage with BsaI exposes complimentary sequences for fragment assembly. Note that the correctly assembled fragments no longer contain BsaI sites and therefore only the plasmids with the correct insertion will remain intact and ready for the last step, which is transformation.
Insert fragments can also be cloned into plasmids and flanked by BsaI sites. During assembly, inserts in vector will ligate to yield the final assembly product which lacks BsaI sites.
Again, only the plasmids with the correct insertion will remain intact and ready for transformation.
For a list of other Type IIS restriction enzymes that can be used for Golden Gate assembly or other applications, please visit www.neb.com/TypeIIsTable.
Online Tools and Resources presentation:
NEB offers many free, easy to use web tools that can help you design your primers for DNA assembly experiments. NEBuilder assembly tool will design your primers for both NEBuilder HiFi DNA Assembly as well as Gibson Assembly.
Simply provide basic information such as the number of fragments you are assembling, and the PCR enzyme you are using. Build your construct by adding fragments to your assembly. The tool provides a list of required primers, recommended annealing temperatures, the final sequence of your assembled construct and a summary of the design.
In conjunction with Benchling, NEB developed the NEB Golden Gate Assembly Tool. The tool automates in silico design of DNA using Golden Gate assembly. Simply select which type IIS restriction enzyme you'd like to use and the number of fragments you're assembling. Provide sequence information for your destination vector and inserts. The tool does the rest.
NEB is committed to providing our customers with quality, no-cost web tools that help to make doing science even easier.
Inside Tech Support No colonies with DNA Assembly
Cathy Shea.:
A common question we get is why am I not seeing colonies when I transform my NEBuilder HiFi DNA Assembly reaction? One of the first things I recommend is to take a look at the competent cells. It's important to use cells that have a high transformation efficiency.
Our cells have a transformation efficiency of greater than 10 x9 colony forming units per microgram. We find that using NEB competent cells often solves the issue of low colony counts. Sub-cloning efficiency cells lower efficiency cells from other vendors and home brewed cells will lead to low colony numbers.
I often recommend a number of products that include competent cells. NEBuilder HiFi DNA Assembly cloning kit includes NEB 5 alpha cells. That's a good choice for routine assemblies. NEBuilder HiFi DNA Assembly bundle for large fragments, includes NEB 10 beta cells. Those are a great choice when your assembly is larger than 15 KB.
Other NEB competent cells also work well with NEBuilder HiFi DNA Assembly. For example, NEB stable cells are a great choice if your assembly includes repeated elements.
You can find additional information about troubleshooting your NEBuilder HiFi DNA Assembly reaction in the manual or by visiting our website.
Deana Martin:
So Chris and Lisa, thanks so much for joining me today.
Chris Teron:
Thanks for having us.
Deana Martin:
It was really fun.
Lisa Maduzia:
It was fun.
Deana Martin:
And always, if you have any suggestions for future episodes, please let us know.
Lisa Maduzia:
Bye.
Related Videos
-
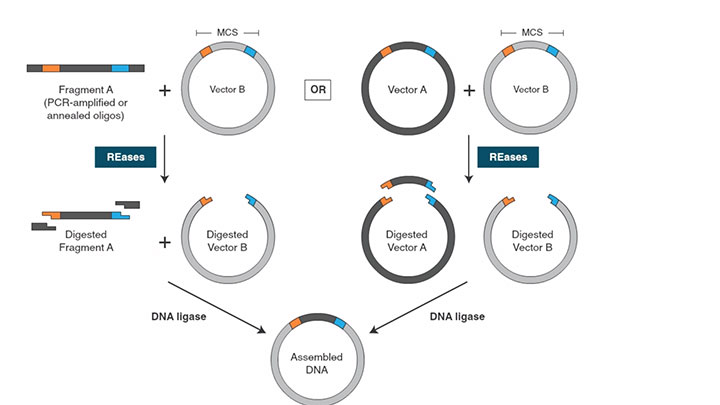
Traditional Cloning Workflow -
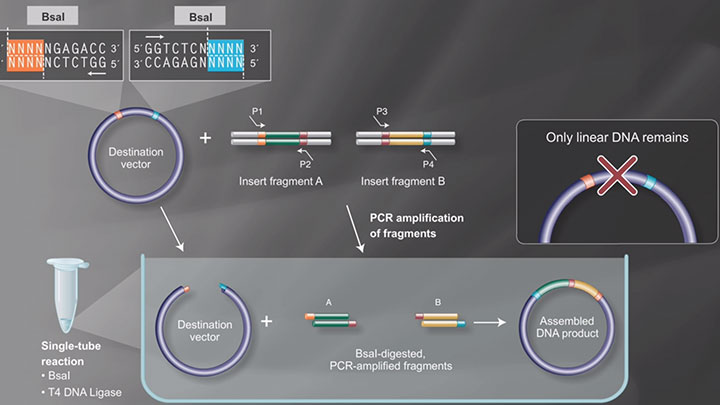
Golden Gate Assembly Workflow -
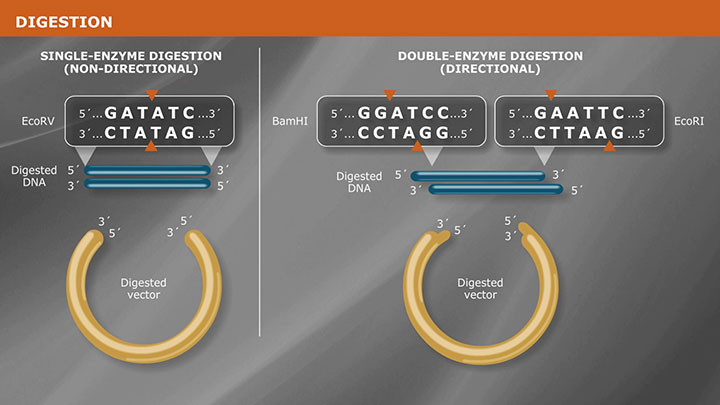
Overview of Traditional Cloning

